|
Use of MASTER to monitor primer-independent and primer-dependent transcription initiation. In primer-dependent transcription initiation, the 3′ nucleotide of a two-, three-, or four-nucleotide RNA primer (di-, tri-, or tetra-nucleotide primer, respectively) base pairs to the template-strand nucleotide in the RNAP active-center P site, and the 5′ nucleotide of the primer base pairs to the template-strand nucleotide in the P−1, P−2, or P−3 site (TSS−1, TSS−2, and TSS−3, respectively Fig. In primer-independent initiation, the initiating entity (typically a nucleoside triphosphate ) base pairs to the template-strand nucleotide in the RNAP active-center P site (TSS Fig. RNAPs can initiate transcription using either a primer-independent or primer-dependent mechanism ( 1 – 10). In transcription initiation, the RNA polymerase (RNAP) holoenzyme binds promoter DNA by making sequence-specific interactions with core promoter elements and unwinds a turn of promoter DNA forming an RNAP-promoter open complex (RPo) containing a single-stranded “transcription bubble.” Next, RNAP selects a transcription start site (TSS) by placing the start-site nucleotide and the next nucleotide of the “template DNA strand” into the RNAP active-center product site (P site) and addition site (A site), respectively, and binding an initiating entity in the RNAP active-center P site ( Fig. Our findings provide a mechanistic and structural description of how TSS-region sequence hard-codes not only the TSS position but also the potential for epitranscriptomic regulation through primer-dependent transcription initiation. coli promoters support the conclusions from MASTER. Results from analysis of a large set of natural, chromosomally encoded E. Biochemical and structure-determination studies show that the base pair (nontemplate-strand base:template-strand base) immediately upstream of the primer binding site (Y:R TSS−2, where R is purine) exerts its effect through the base on the DNA template strand (R TSS−2) through interchain base stacking with the RNA primer. The results yield a consensus sequence for primer-dependent initiation, Y TSS−2N TSS−1N TSSW TSS+1, where TSS is the transcription start site, N TSS−1N TSS is the primer binding site, Y is pyrimidine, and W is A or T. coli involves any of the 16 possible dinucleotide primers and depends on promoter sequences in, upstream, and downstream of the primer binding site. The results show primer-dependent initiation in E.  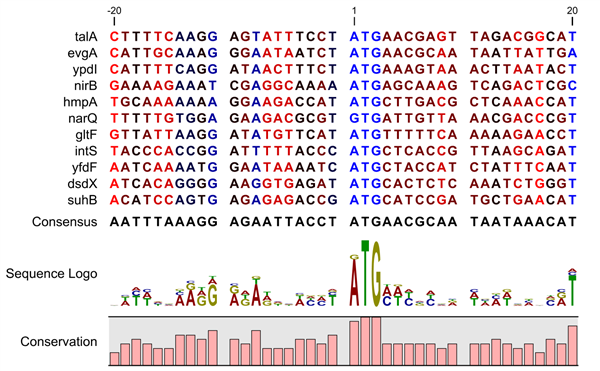
In Escherichia coli cells, RNAs with a 5′-end hydroxyl are generated by use of dinucleotide RNAs as primers for transcription initiation, “primer-dependent initiation.” Here, we use massively systematic transcript end readout (MASTER) to detect and quantify RNA 5′-ends generated by primer-dependent initiation for ∼4 10 (∼1,000,000) promoter sequences in E. In transcription initiation, a large inventory of substrates compete with nucleoside triphosphates for use as initiating entities, providing an ab initio mechanism for altering the RNA 5′-end. Chemical modifications of RNA 5′-ends enable “epitranscriptomic” regulation, influencing multiple aspects of RNA fate.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
